 PDF(4747 KB)
PDF(4747 KB)


基于无人机高光谱数据和3D-2D-CNN的天然次生林主要树种分类研究
李昊, 全迎, 刘建阳, 卞少杰, 王斌, 李明泽
南京林业大学学报(自然科学版) ›› 2026, Vol. 50 ›› Issue (2) : 9-18.
 PDF(4747 KB)
PDF(4747 KB)
 PDF(4747 KB)
PDF(4747 KB)
基于无人机高光谱数据和3D-2D-CNN的天然次生林主要树种分类研究
Discriminate dominant tree species in natural secondary forests from UAV hyperspectral images using a hybrid 3D-2D convolutional neural network
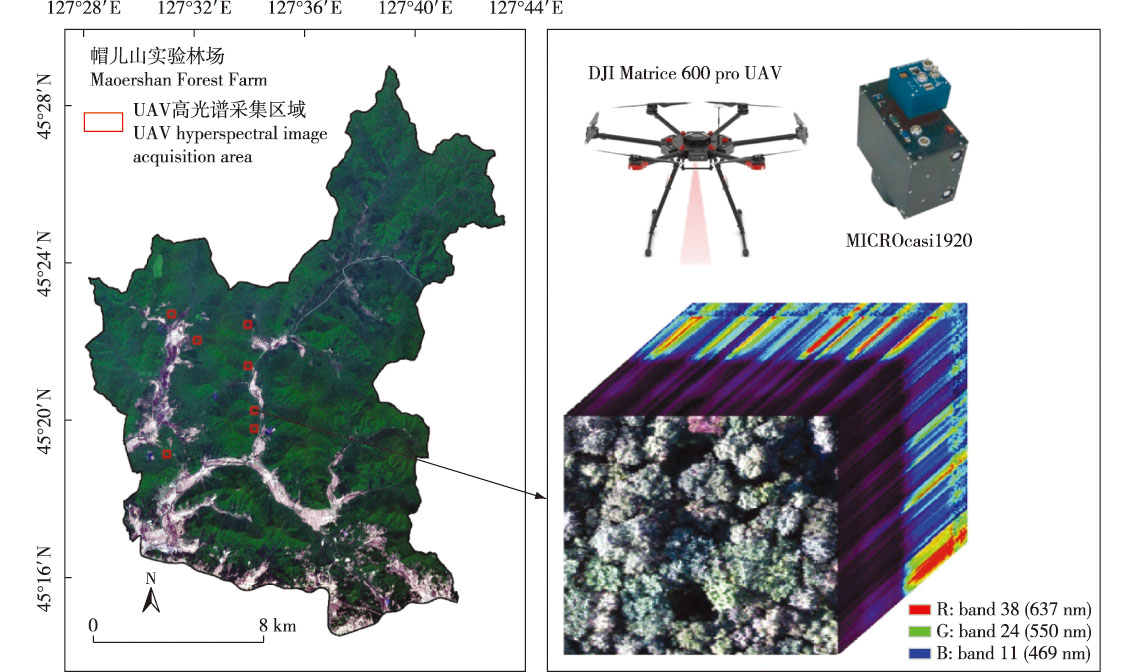
【目的】为了提高我国东北典型天然次生林主要树种分类的准确性,提出一种针对无人机高光谱图像的卷积神经网络(convolution neural network,CNN)树种分类框架。【方法】以东北林业大学帽儿山实验林场天然次生林水曲柳(Fraxinus mandshurica)、胡桃楸(Juglans mandshurica)、榆树(Ulmus sp.)和白桦(Betula platyphylla)的4个主要树种为研究对象,采用新型无人机载高光谱成像仪对7个不同区域进行影像采集。以地面实测数据为参考构建不同树冠尺寸的单木数据集,并以7∶3的比例划分训练集和测试集,构建了包含三维卷积层和二维卷积层的3D-2D-CNN模型,通过3D卷积层提取光谱-空间耦合特征,2D卷积层提取细节特征,从而有效增强模型对数据整体特征的学习能力;将该模型与2D-CNN、3D-CNN以及基于特征筛选的机器学习模型[随机森林(random forest,RF)、支持向量机(support vector machine,SVM)和梯度增强机(gradient boosting machine,GBM)]进行对比实验。此外,采用逐波段逐步移除方法分析了波段的重要性,并探讨了模型对光谱特征的敏感性。【结果】构建的3D-2D-CNN分类模型对研究区4个主要树种的分类准确率达到了87%,F1分数为0.86,相较其他对比算法,总体精度提高了5%~6%。波段重要性分析表明,近红外波段对分类结果影响显著。【结论】基于高光谱图像的3D-2D-CNN模型通过有效结合光谱与空间信息,显著提升了对天然次生林树种分类的准确性,比传统分类方法表现优越,可为森林资源管理和生态系统的遥感监测提供技术支持。
【Objective】This study aims to improve the classification accuracy of dominant tree species in typical natural secondary forests in northeast China, a convolutional neural network (CNN)-based framework for tree species classification using UAV hyperspectral images was proposed.【Method】Four dominant tree species—Fraxinus mandshurica, Juglans mandshurica, Ulmus sp., and Betula platyphylla—from the Maoershan Experimental Forest Farm of Northeast Forestry University were studied. Hyperspectral images of seven different regions were acquired using a novel UAV-mounted hyperspectral imager. A single-tree dataset with varying crown sizes was constructed using ground-measured data, divided into training and test sets at a 7∶3 ratio. A hybrid 3D-2D-CNN model integrating 3D and 2D convolutional layers was developed: 3D convolutional layers extracted spectral-spatial coupled features, while 2D layers captured detailed spatial features, enhancing the model’s holistic learning capability. The model was compared with 2D-CNN, 3D-CNN, and feature-selection-based machine learning models (random forest (RF), support vector machine (SVM), and gradient boosting machine (GBM)). Additionally, the band importance was analyzed using a progressive band removal method, and spectral feature sensitivity was investigated.【Result】The proposed 3D-2D-CNN model achieved a classification accuracy of 87% and an F1 score of 0.86 for the four tree species, outperforming other algorithms with an overall accuracy improvement of 5%-6%. Band importance analysis highlighted the significant contribution of the near-infrared band classification.【Conclusion】The 3D-2D-CNN model, by effectively integrating spectral and spatial information, significantly enhanced the classification performance of natural secondary forest tree species compared to traditional methods. This approach provides technical support for forest resource management and ecosystem monitoring via remote sensing.
天然次生林 / 树种分类 / 高光谱图像 / 3D-2D-CNN模型 / 卷积神经网络 / 机器学习 / 波段重要性分析
natural secondary forest / tree species classification / hyperspectral image / 3D-2D-CNN model / convolutional neural network / machine learning / band importance analysis
| [1] |
|
| [2] |
吴见, 彭道黎. 高光谱遥感林业信息提取技术研究进展[J]. 光谱学与光谱分析, 2011, 31(9):2305-2312.
|
| [3] |
|
| [4] |
|
| [5] |
|
| [6] |
|
| [7] |
|
| [8] |
|
| [9] |
|
| [10] |
|
| [11] |
|
| [12] |
|
| [13] |
|
| [14] |
|
| [15] |
|
| [16] |
|
| [17] |
|
| [18] |
|
| [19] |
|
| [20] |
|
| [21] |
|
| [22] |
|
| [23] |
|
| [24] |
|
| [25] |
|
| [26] |
|
| [27] |
|
| [28] |
|
| [29] |
|
| [30] |
|
| [31] |
|
| [32] |
|
| [33] |
|
| [34] |
|
| [35] |
|
| [36] |
|
| [37] |
|
| [38] |
|
| [39] |
|
| [40] |
|
| [41] |
|
| [42] |
|
| [43] |
全迎. 融合无人机LiDAR和高光谱特征的天然次生林树种及林分类型识别[D]. 哈尔滨: 东北林业大学, 2023.
|
| [44] |
赵颖慧, 张大力, 甄贞. 基于非参数分类算法和多源遥感数据的单木树种分类[J]. 南京林业大学学报(自然科学版), 2019, 43(5):103-112.
|
| [45] |
|
| [46] |
|
/
| 〈 |
|
〉 |