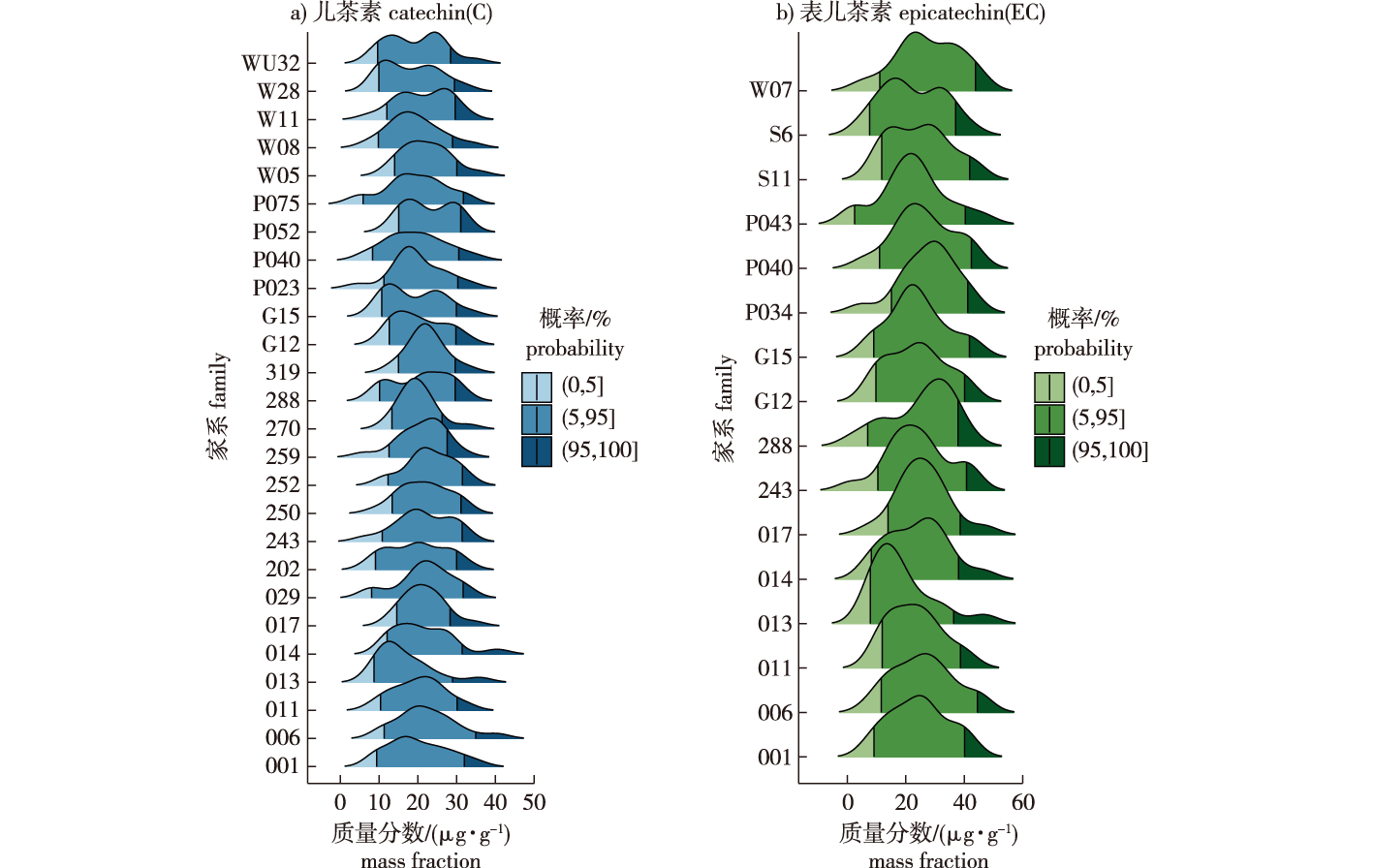
【Objective】Pinus taeda (loblolly pine) stands as a globally significant industrial timber species, where genetic improvement efforts have historically centered on growth traits. Given the pivotal role of catechins within the phenylpropanoid metabolic pathway, their potent capacity for scavenging reactive oxygen species (ROS), their contribution to enhanced plant stress resistance, and their substantial application potential in pharmaceutical and food industries, the screening of loblolly pine germplasm resources exhibiting high catechin content is of considerable scientific and practical importance. This study aims to evaluate the genetic variation of catechin and epicatechin contents among and within families of Pinus taeda (loblolly pine). Based on the genetic variation characteristics, superior families and elite individual trees are identified, providing materials and theoretical references for the cultivation of high-catechin Pinus taeda plantations and the selective breeding of fine varieties.【Method】The trial was conducted within a plantation managed by the Forestry Science Research Institute of Yingde City, Guangdong Province, China (113°45' E,24°15' N). The plant material comprised open-pollinated progenies from a 1.5-generation seed orchard of loblolly pine, established in spring 2015. A randomized complete block design was implemented, consisting of eight blocks and 54 families. Each plot contained five trees planted in a single row, with a spacing of 3 m × 3 m, resulting in a total of 864 individual trees sampled. Needle samples were collected, and their diffuse reflectance spectra were acquired using a Perten DA7200 near-infrared (NIR) spectrometer across a wavelength range of 950-1 650 nm, at a resolution of 5 nm, and a scanning speed of approximately 100 spectra per second. Each sample was repacked and scanned three times, yielding a total of 9 spectra per sample. Utilizing a previously established and validated NIR prediction model for needle catechin and epicatechin contents (calibration set: R2 = 0.92, RMSEC = 1.53 μg/g; validation set: R2 = 0.88, RMSEP = 1.84 μg/g, RPD = 3.1), spectral data preprocessing and quantitative analysis were performed on Unscrambler X 10.4 software platform. Preprocessing involved Standard Normal Variate (SNV) transformation combined with Savitzky-Golay first-derivative smoothing for noise reduction. Data visualization and fundamental statistical analysis were conducted using R software (version 4.2.3). Genetic analyses employed the ASReml-R V4 software platform to fit mixed linear models. Best Linear Unbiased Prediction (BLUP) was used to estimate family effects and individual tree breeding values. Key genetic parameters, including family heritability, individual heritability, and within-family heritability, were calculated. Analysis of variance (ANOVA) was performed to assess the significance of effects due to family, block, and their interaction.【Result】Genetic variance analysis revealed that the contents of both catechin (C) and epicatechin (EC) in loblolly pine needles exhibited highly significant variation (P < 0.01) among families and for the family-by-block interaction effect, indicating substantial exploitable genetic variation and significant selection potential. Estimates of genetic parameters were as follows: family heritability was 0.376 ± 0.131 for C and 0.292 ± 0.147 for EC; Individual heritability was 0.172 ± 0.091 for C and 0.124 ± 0.084 for EC; Within-family heritability was 0.146 ± 0.080 for C and 0.104 ± 0.073 for EC. The performance of elite individuals selected via the three methods was characterized: (1) Direct breeding value selection. Selected individuals exhibited C breeding values ranging from 2.034 2-2.886 4 and EC breeding values ranging from 2.409 4-3.268 0. This method yielded the highest individual breeding values but resulted in selections concentrated within fewer families. (2) Selection controlling family co-ancestry. For C, eight families were selected (two trees per family), with breeding values ranging from 1.793 1-2.886 4. For EC, nine families were selected (two trees per family), with breeding values ranging from 2.262 8-3.268 0. This strategy effectively combined genetic gain at both the family and individual levels. (3) Combined selection. For C, this strategy selected five families (two trees per family) and seven families (one tree per family), yielding genetic gains ranging from 1.390 9-2.886 4. For EC, it selected five families (two trees per family) and seven families (one tree per family), yielding genetic gains ranging from 1.665 6-3.267 9.【Conclusion】This study conclusively demonstrates the existence of substantial and exploitable genetic variation in catechin and epicatechin contents within loblolly pine needles. Through systematic genetic evaluation and multi-strategy selection, superior families and elite individual trees exhibiting high breeding values and predicted genetic gains for both C and EC were successfully identified. Targeting catechin contents as an independent and crucial breeding objective, distinct from traditional growth traits, represents a significant paradigm shift. The selected elite germplasm provides invaluable genetic resources for advancing high-generation genetic improvement programs in loblolly pine. Moreover, these materials can be directly utilized as superior propagation stock for establishing plantations specifically dedicated to high catechin production. This research holds considerable theoretical and practical significance for propelling the selective breeding of loblolly pine towards enhanced functional compound utilization and value-added applications.
 PDF(1937 KB)
PDF(1937 KB)


 PDF(1937 KB)
PDF(1937 KB)
 PDF(1937 KB)
PDF(1937 KB)