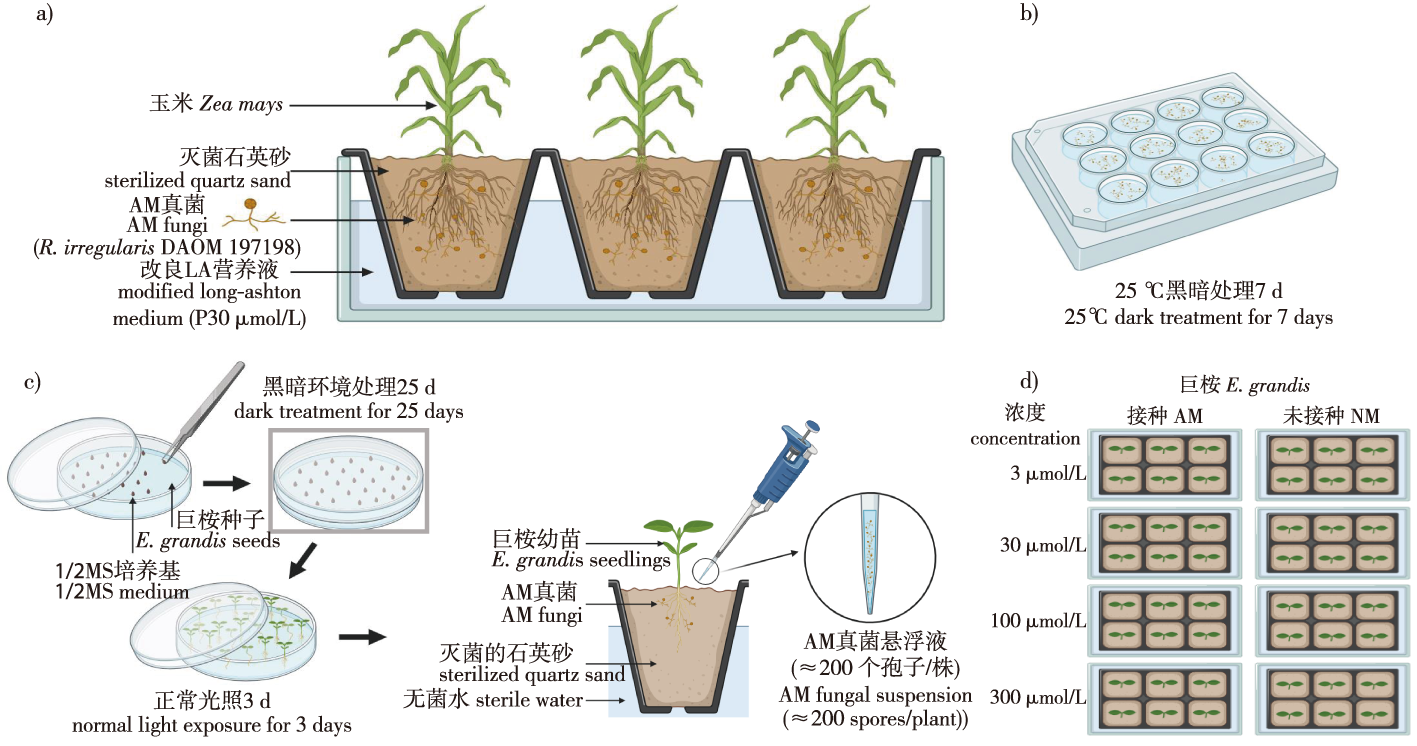
【Objective】Low phosphorus stress is one of the primary factors limiting the growth and development of eucalyptus (Eucalyptus grandis). Arbuscular mycorrhiza (AM) fungi, through their symbiotic relationship with plants, are known to enhance phosphorus absorption, particularly under low phosphorus conditions. This mutualistic association plays a crucial role in alleviating phosphorus limitation in plants by increasing the surface area for nutrient uptake and facilitating the transport of phosphorus from soil to plant. To investigate the effect of AM fungi on the expression of phosphorus (P) starvation response genes in E. grandis under low-P conditions, this study analyzed the expression levels of SPX and PHR genes, which are involved in P starvation response, in E. grandis seedlings inoculated with AM fungi across different phosphorus concentrations. Specifically, the study focused on the role of AM fungi in modulating the expression of key genes involved in the phosphate starvation response, namely, PHR and SPX genes. 【Method】 E. grandis seedlings were inoculated with the AM fungus Rhizophagus irregularis under varying phosphorus supply levels. Four phosphorus treatments were applied: extremely low (3 μmol/L NaH2PO4), low (30 μmol/L), medium (100 μmol/L) and high (300 μmol/L). The seedlings were grown under controlled conditions, and various plant physiological parameters, including plant height, root length and biomass, were measured to assess the impact of the phosphorus supply and AM fungi inoculation on growth. The gene expression levels of phosphate starvation response genes, particularly EgPHR2 and EgSPX2, were analyzed through quantitative PCR to identify key genes involved in the phosphorus acquisition pathway and their regulation under different phosphorus conditions.【Result】Mycorrhizal inoculation had a significant impact on the colonization rates of E. grandis roots, especially under low phosphorus conditions. The mycorrhizal colonization rates for seedlings treated with extremely low (3 μmol/L NaH2PO4) and low (30 μmol/L NaH2PO4) phosphorus were 82.61% and 67.53%, respectively, those were significantly higher than the colonization rates under medium (100 μmol/L NaH2PO4, 38.60%) and high phosphorus (300 μmol/L NaH2PO4, 38.64%) treatments. In addition to improved colonization, AM fungi inoculation also led to significant increases in plant height, root length, and biomass, particularly under phosphorus-limited conditions. Bioinformatics analysis of the E. grandis genome identified two PHR genes and six SPX genes that are likely involved in the phosphate starvation response. The expression of EgPHR2 and EgSPX2 in AM-inoculated seedlings was significantly higher compared to non-inoculated controls. Specifically, the expression of EgPHR2 increased by 273.64%, 294.67%, and 698.15% at 30 μmol/L, 100 μmol/L and 300 μmol/L phosphorus, respectively. Similarly, EgSPX2 expression showed substantial increases of 2 517.15%, 606.40% and 923.13% under the same phosphorus conditions, indicating a strong upregulation of these genes in response to phosphorus limitation.【Conclusion】The findings of this study suggest that EgPHR2 and EgSPX2 are specifically involved in the expression is closely linked to the colonization of E. grandis roots by AM fungi. The expression of EgPHR2 was particularly elevated under high phosphorus conditions, while EgSPX2 showed higher expression under low phosphorus conditions, highlighting their distinct roles in phosphorus homeostasis. The results also demonstrate that AM fungi play a crucial role in E. grandis, particularly under phosphorus-deficient conditions, by modulating the expression of key genes involved in phosphorus acquisition and signaling pathways.
 PDF(3898 KB)
PDF(3898 KB)


 PDF(3898 KB)
PDF(3898 KB)
 PDF(3898 KB)
PDF(3898 KB)