 PDF(2727 KB)
PDF(2727 KB)


Bioinformatics analysis of the CcdC protein of the DUF1453 family in Bacillus velezensis FZB42 and its effects on growth and development
LI Meiju, SHEN Zizhu, GUAN Chenyun, FAN Ben, ZHAO Yinjuan
Journal of Nanjing Forestry University (Natural Sciences Edition) ›› 2025, Vol. 49 ›› Issue (6) : 247-254.
 PDF(2727 KB)
PDF(2727 KB)
 PDF(2727 KB)
PDF(2727 KB)
Bioinformatics analysis of the CcdC protein of the DUF1453 family in Bacillus velezensis FZB42 and its effects on growth and development
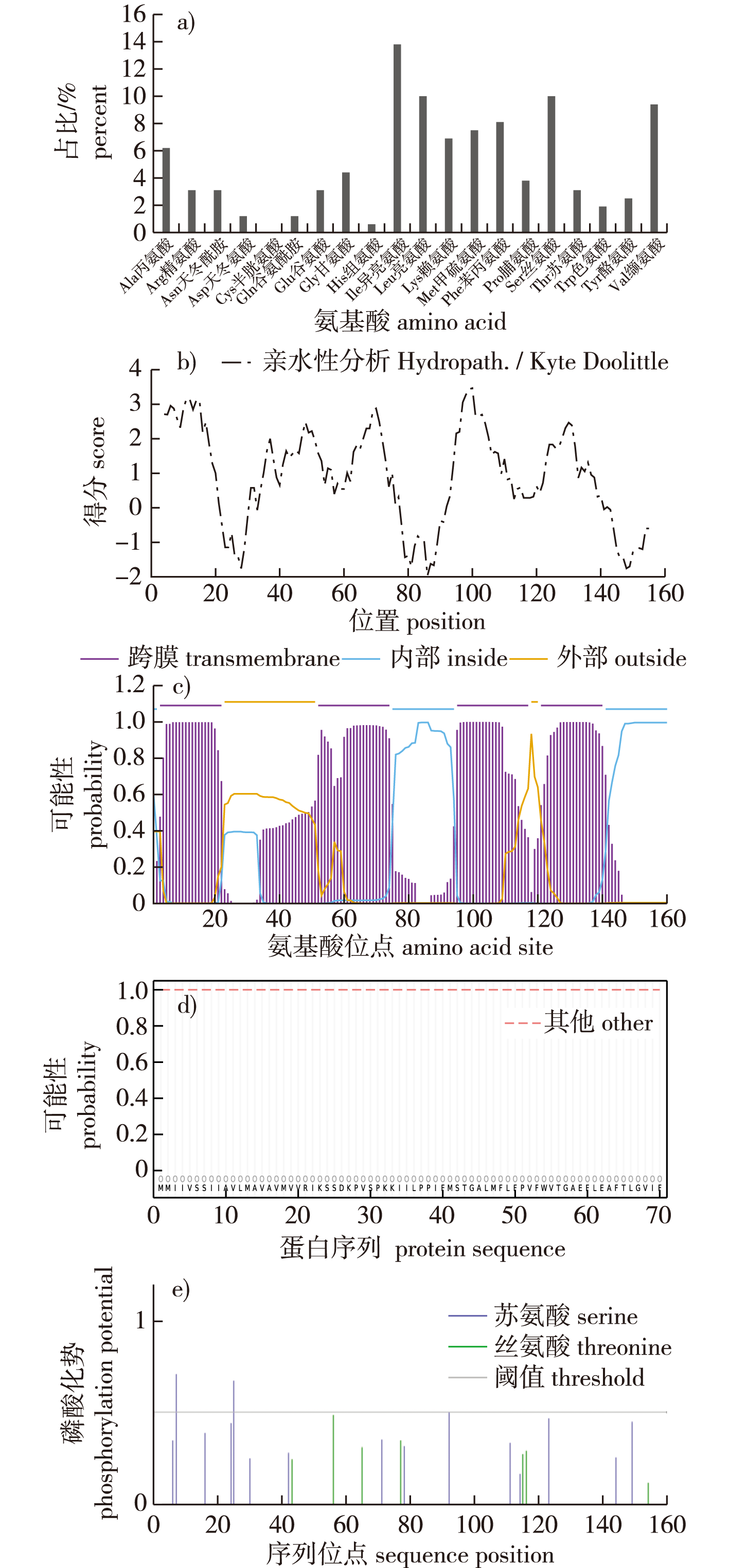
【Objective】The CcdC protein in Bacillus velezensis FZB42 contains a DUF1453 domain with an unknown structure and function. This study aims to predict the biological properties of CcdC using bioinformatics methods and to investigate its function by constructing a ccdC knockout strain, thereby laying the groundwork for elucidating the structure and function of the DUF1453 domain.【Method】Bioinformatics online tools were employed to predict the amino acid composition, physicochemical properties, hydrophobicity, transmembrane regions, and subcellular localization of the CcdC protein. A phylogenetic tree analysis of CcdC proteins from different bacterial species was also conducted. A ccdC knockout strain of B. velezensis FZB42 was constructed, and the growth of the wild-type FZB42 strain and the ccdC knockout strain was compared in both liquid and solid LB media. Additionally, spore formation of both strains was assessed in DSM medium.【Result】The CcdC protein consisted of 160 amino acids and was a basic, stable, hydrophobic transmembrane protein located in the cell membrane, lacking a signal peptide. Secondary structure analysis revealed that the CcdC protein was predominantly α-helical. Tertiary structure analysis indicated that no structurally similar proteins to CcdC existed in current databases. Colony PCR confirmed the successful construction of the ccdC knockout strain. Comparative analysis of growth and spore formation between the wild-type FZB42 and the ccdC knockout strain demonstrated that the deletion of the ccdC gene significantly affected bacterial growth and spore formation.【Conclusion】Bioinformatics analysis tentatively identified CcdC as a membrane protein, with predicted phosphorylation sites suggesting its potential involvement in signal transduction functions. Interaction network analysis indicated that CcdC protein may be related to growth and development. The construction of the knockout strain confirmed that the deletion of the ccdC gene impair bacterial growth and development, providing a foundation for further research into the biological function of the DUF1453 domain and the regulatory role of CcdC in the growth and development of B. velezensis FZB42.
DUF1453 domain / CcdC protein / bioinformatics analysis / growth and development / Bacillus velezensis
| [1] |
|
| [2] |
|
| [3] |
|
| [4] |
|
| [5] |
|
| [6] |
|
| [7] |
|
| [8] |
|
| [9] |
|
| [10] |
|
| [11] |
|
| [12] |
|
| [13] |
|
| [14] |
|
| [15] |
|
| [16] |
|
| [17] |
|
| [18] |
|
| [19] |
|
| [20] |
孙柯, 赵学亮, 白丽艳, 等. 布鲁氏杆菌S2株DUF2326基因克隆与生物信息学分析[J]. 黑龙江畜牧兽医, 2020(5):9-14,148.
|
| [21] |
|
| [22] |
|
| [23] |
|
| [24] |
|
| [25] |
|
| [26] |
|
/
| 〈 |
|
〉 |