 PDF(4333 KB)
PDF(4333 KB)


Analysis of rhizosphere soil bacterial community characteristics and functional bacterial excavation of Tamarix austromongolica in Qinghai Province
ZHOU Yanxu, LI Qiang, LIU Lei, WANG Xiangfu, SUN Qiwu, YANG Fu, HOU Lingyu, LIU Yong, WU Lianghou, ZHOU Hongquan, WANG Baorong
Journal of Nanjing Forestry University (Natural Sciences Edition) ›› 2026, Vol. 50 ›› Issue (2) : 236-244.
 PDF(4333 KB)
PDF(4333 KB)
 PDF(4333 KB)
PDF(4333 KB)
Analysis of rhizosphere soil bacterial community characteristics and functional bacterial excavation of Tamarix austromongolica in Qinghai Province
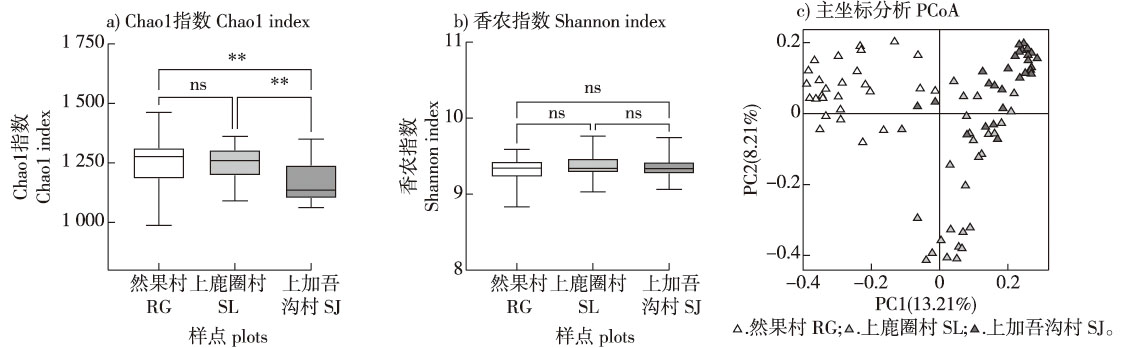
【Objective】This study systematically analyzed the diversity, community composition, and growth-promoting functions of rhizosphere soil bacteria in Tamarix austromongolica from Qinghai Province. The research aims to provide a basis for the conservation of this tree species and the development of its indigenous microbial resources.【Method】Using the rhizosphere soil of T. austromongolica in three conservation areas, Ranguo Village (RG), Shanglujuan Village (SL), and Shangjiawugou Village (SJ) in Hainan Zang Autonomous Prefecture, Qinghai Province as the research object, the bacterial diversity and community composition of the rhizosphere soil of T. austromongolica were analyzed based on high-throughput sequencing, rhizosphere soil bacteria were isolated using the traditional dilution culture method, and functional evaluation was performed with functional plates and specific color changes.【Result】There are significant differences in the Alpha and Beta diversity of rhizosphere soil bacteria in T. austromongolica from Ranguo Village (RG) compared to Shanglujuan Village (SL) and Shangjiawugou Village (SJ). However, the dominant bacterial phyla across all three conservation areas are Proteobacteria, Acidobacteriota, and Actinomycetota. Pseudomonas, unclassified_bacteria, and unclassified_Gemmatimonadaceae are the biomarker for the different conservation areas, respectively. A total of 55 strains are isolated from the rhizosphere soil of T. austromongolica across different conservation areas, among which two strains are potential new species, and 11 strains belong to the core genera of Streptomyces, Sphingomonas, Lysobacter and Devosia. Twenty-seven strains exhibit potential growth-promoting functions, with 12 strains processing inorganic phosphorus-dissolving ability, eight strains demonstrating organic phosphorus-mineralizing ability, eight strains showing potassium-dissolving ability, and 15 strains capable of IAA-producing ability. Among there, the strain Oceanbacillus sojae RGGN1 isolated from RG possessed the functions of inorganic phosphorus-dissolving, organic phosphorus-mineralizing, potassium dissolving and IAA-producing ability. Simultaneously possess three of these functions.【Conclusion】There are certain differences in the rhizosphere soil bacterial diversity and community composition of T. austromongolica in different conservation areas. Using culturable methods, 11 core groups were identified, and two potential new species along with 27 strains exhibiting potential growth-promoting functions were obtained. This work enriches the resource base of high-quality functional bacterial strains with regional characteristics and independent intellectual property rights.
Tamarix austromongolica / plant growth-promoting rhizobacteria / community structure / high-throughput sequencing / isolation and cultivation / function bacterial assessment / Qinghai
| [1] |
|
| [2] |
方欧娅, 贾恒锋, 邱红岩, 等. 青海省同德县乔木状甘蒙柽柳的年龄及其生长对环境的响应[J]. 植物生态学报, 2017, 41(7):738-748.
|
| [3] |
|
| [4] |
孔亚丽, 朱春权, 曹小闯, 等. 土壤微生物介导植物抗盐性机理的研究进展[J]. 中国农业科学, 2021, 54(10):2073-2083.
|
| [5] |
宋健, 刘伟峰, 魏喜喜, 等. 枣专用微生物菌剂对干旱区骏枣园土壤养分及土壤酶活性的影响[J]. 西南农业学报, 2021, 34(7):1472-1479.
|
| [6] |
刘心葵, 武崇高, 刘雪峰, 等. 不同地区樟子松枯梢病林下土壤真菌多样性[J]. 森林工程, 2024, 40 (6):41-52.
|
| [7] |
刘浩南, 丁珊珊, 包小春, 等. 3株解磷菌的解磷机理及其对杉木幼苗的促生效应[J]. 生态与农村环境学报, 2025, 41(3):378-388.
|
| [8] |
|
| [9] |
|
| [10] |
申建波, 白洋, 韦中, 等. 根际生命共同体:协调资源、环境和粮食安全的学术思路与交叉创新[J]. 土壤学报, 2021, 58(4):805-813.
|
| [11] |
|
| [12] |
董醇波, 张芝元, 韩燕峰, 等. 核心微生物组的研究及利用现状[J]. 菌物学报, 2019, 38(1):1-10.
|
| [13] |
张小霞, 陈筱玥, 王秋云, 等. 柽柳根际一株盐单胞菌Bachu 26的分离、鉴定及其盐胁迫下的促生作用研究[J]. 微生物学报, 2024, 64(2):607-622.
|
| [14] |
潘雅清. 荒漠草原盐生植物—微生物互馈效应研究[D]. 银川: 宁夏大学, 2022.
|
| [15] |
陈乙煌, 邢利, 东珍珍, 等. 柽柳根际土壤放线菌的分离及拮抗活性筛选[J]. 新疆农业科学, 2023, 60(7):1790-1797.
|
| [16] |
|
| [17] |
杨芾, 王辉, 王琴, 等. 浙江地区毛竹林根际土壤促生细菌筛选及促生效应研究[J]. 南京林业大学学报(自然科学版), 2024,1-10.
|
| [18] |
|
| [19] |
|
| [20] |
王阳, 王伟, 姜静, 等. 转基因小黑杨根际土壤微生物群落特征研究[J]. 南京林业大学学报(自然科学版), 2023, 47(1):199-208.
|
| [21] |
|
| [22] |
HUHE,
|
| [23] |
|
| [24] |
|
| [25] |
丁成翔. 青藏高原高寒草原土壤微生物对不同放牧方式的响应[D]. 西宁: 青海大学, 2020.
|
| [26] |
|
| [27] |
|
| [28] |
王改萍, 阿依古丽·托乎提, 王茹, 等. 新疆乌尔禾地区盐渍土壤耐盐细菌多样性与群落结构研究[J]. 微生物学杂志, 2021, 41(2):17-26.
|
| [29] |
代金霞, 田平雅, 张莹, 等. 银北盐渍化土壤中6种耐盐植物根际细菌群落结构及其多样性[J]. 生态学报, 2019, 39(8):2705-2714.
|
| [30] |
周楠, 姜成英, 刘双江. 从环境中分离培养微生物:培养基营养水平至关重要[J]. 微生物学通报, 2016, 43(5):1075-1081.
|
| [31] |
贾向飞, 曹琨, 吴斌, 等. 产耐酸纤维素酶放线菌的筛选、产酶优化及酶学性质[J]. 生物加工过程, 2025, 23(3):252-261.
|
| [32] |
张嘉馨, 骆新荣, 刘占文, 等. 塔克拉玛干沙漠来源产氨基糖苷类次级代谢产物潜力链霉菌的筛选[J]. 塔里木大学学报, 2024, 36(1):13-21.
|
| [33] |
刘辉, 韦璐璐, 朱龙发, 等. 鞘氨醇单胞菌的研究进展[J]. 微生物学通报, 2023, 50(6):2738-2752.
|
| [34] |
|
| [35] |
|
| [36] |
周泽, 姚拓, 史潭梅, 等. 菌剂对高寒地区土壤微生物群落结构及固氮菌群的影响[J]. 草地学报, 2022, 30 (10):2609-2616.
|
| [37] |
|
| [38] |
|
| [39] |
|
/
| 〈 |
|
〉 |